...
The Hierarchical Editing Language for Macromolecules was designed to create a single notation that can encode the structure of complex biomolecules including diverse polymers, non-natural monomers and complex attachment points.
...
HELM was first conceived at Pfizer in the summer of 2008 to support the Pfizer oligonucleotide therapeutic unit and molecules were first registered into the Pfizer corporate database using HELM in December 2008.
How it works
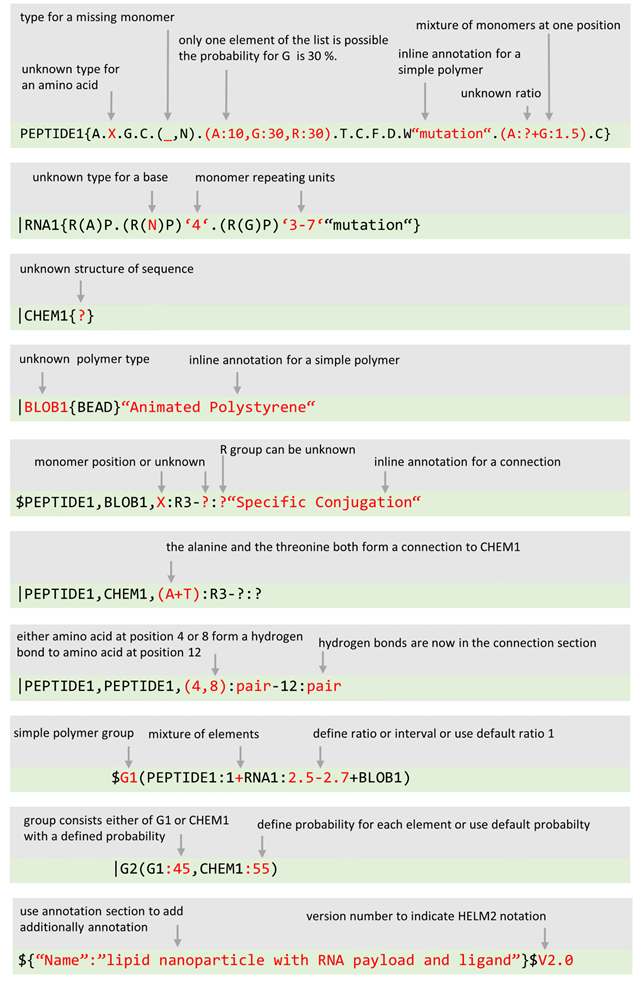
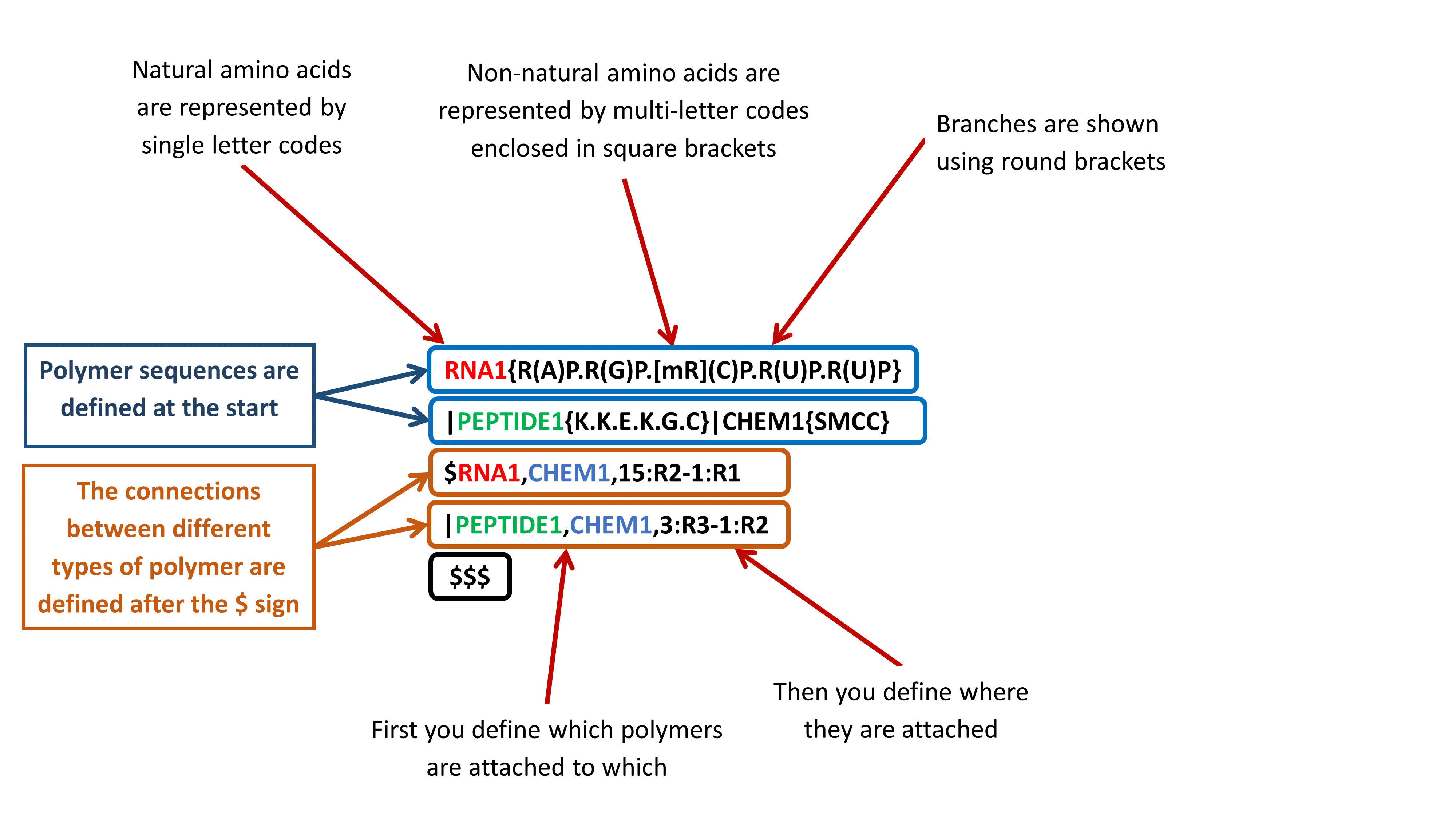
HELM contains multiple levels of information:
...
There are 2 major releases of HELM.
HELM1 | HELM 2 |
|---|---|
Covers different types of macromolecule : DNA/RNA, PEPTIDE and CHEM polymer types Allows non-natural monomers : Monomers are defined by the HELM author, so there is no limit on the monomers that can be included in your molecule. Is portable xHELM allows you to ‘bundle’ all the monomer information with your molecule definition into a single package that can be used to transfer information outside your organisation. | Adds the ability to define
|
Specification
The HELM 2.04 specification is available below.
...