The Hierarchical Editing Language for Macromolecules was designed to create a single notation that can encode the structure of complex biomolecules including diverse polymers, non-natural monomers and complex attachment points.

HELM was first conceived at Pfizer in the summer of 2008 to support the Pfizer oligonucleotide therapeutic unit and molecules were first registered into the Pfizer corporate database using HELM in December 2008.
HELM contains multiple levels of information:

The flexibility allows you to define molecules like this which include a oligonucleotide connected to a small molecule (SMCC) connected to a peptide.
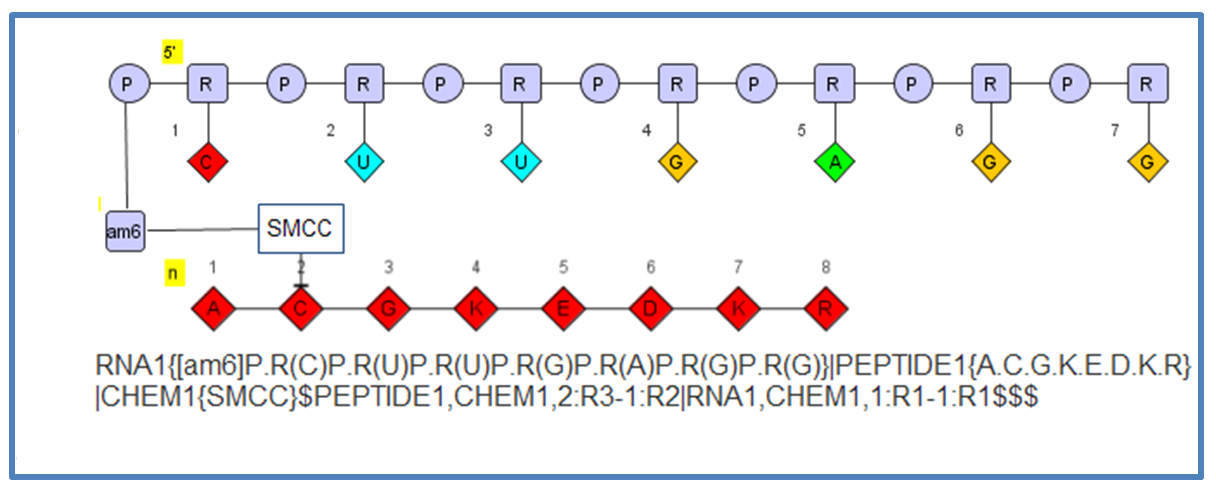
HELM1:
Covers different types of macromolecule
Allows non-natural monomers
Is portable
HELM 2
Adds the ability to define
Specification
The HELM 2.04 specification is available below.
There are minor changes in the 2.04 update. Specifically:
Test set
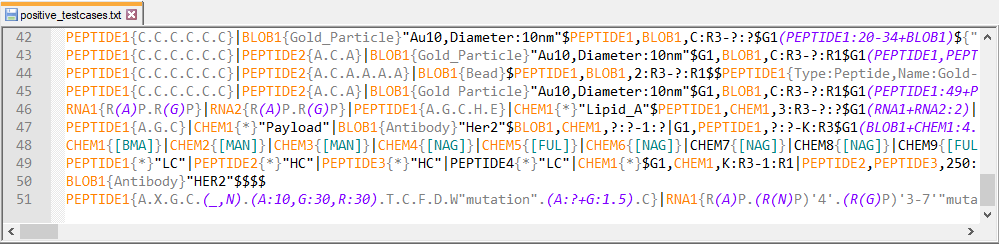
The team have compiled a set of around 150 structures that illustrate the full range of structures that can be encoded in HELM.This is included as a resource for anyone who wants to implement their own HELM tools rather than use the open source toolkit.
Peptide Monomer Guidelines
We have developed recommendations for creating and naming a monomer set for peptides.
Nucleotide Monomer Guidelines
More Information
An academic overview of HELM and its origins
HELM 1.0 overview
xHELM overview
HELM 2.0 quick reference diagram

Notepad++ highlighting definition file
If you create HELM manually it can be helpful to use an editor that will highlight different parts of the string.
To use this language definition file go to [Language] -> [Define you language...]. Here you have to click on [Import...] and load the xml file. After closing the dialog, you have to restart Notepad++. Now you can assign the language to you current open file via [Language] -> [HELM2].