Fragmentation - or polymers to monomers
Many organisations have macromolecule information in a variety of formats. Converting HELM to an atom/bond molecular representation is a relatively straightforward process since the monomers and attachment points between monomers are defined, so assembly is straightforward.
Converting an atom/bond representation to HELM is more complex, since you have to decide what the monomers are.
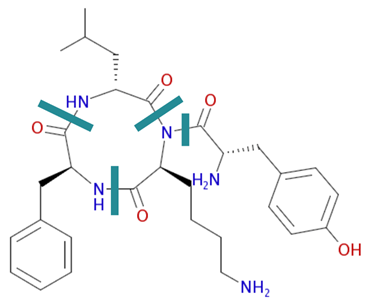
Identifying monomers is not always straightforward. There are decisions about where the backbone actually is which is not always straightforward. See this cyclic peptide as an example:
There are some tools that can help fragment peptides, however the groups that developed them would be the first to say that they are not perfect. We would encourage more work in this area.
Free to use toolsets
The HELM toolkit contains a (very basic) fragmenter, and the EBI has an internal tool, details of which would be available from them on request.